Metabolomics is the study of the small molecules, or metabolites, that drive life's processes. It provides a powerful way to understand the complex link between a living system's chemistry and its observable traits. One of the great promises of this field is the discovery of biomarkers, molecules that can signal health, disease, or response to treatment, offering incredible value for scientific discovery [3].
However, the journey from a biological sample to a clear insight faces a significant hurdle. A recent and sobering meta-analysis of 244 clinical studies concluded that a staggering 85% of proposed biomarkers are simply statistical noise [1]. This finding points to a reproducibility crisis, suggesting that while we are getting better at measuring metabolites, the field of metabolomics analysis needs to improve how we find the signals that truly matter.
The Search for a True Signal: Why Are Studies Struggling?
If so many findings are statistical noise, what is the underlying cause?
A recent study gives us a huge clue. Bifarin et al. (2025) mapped the entire metabolomics research landscape from the past two decades, analyzing over 80,000 publications. Their work revealed a critical fact: the vast majority of studies are too small to find a real signal. Based on the 2,613 studies that mentioned sample size in their abstract, a massive 70.8% used fewer than 50 samples [2].
This presents a fundamental challenge. Small studies often lack the necessary statistical power to distinguish a true biological signal from random variation. As highlighted by Xu & Goodacre (2025), this type of "short fat data", where the number of measured metabolites far exceeds the number of samples, is highly susceptible to statistical caveats. Without rigorous validation, powerful statistical models can easily find patterns in random noise, leading to over-optimistic and irreproducible results [11].
This creates a major bottleneck for the entire field, but it also highlights a clear path forward.
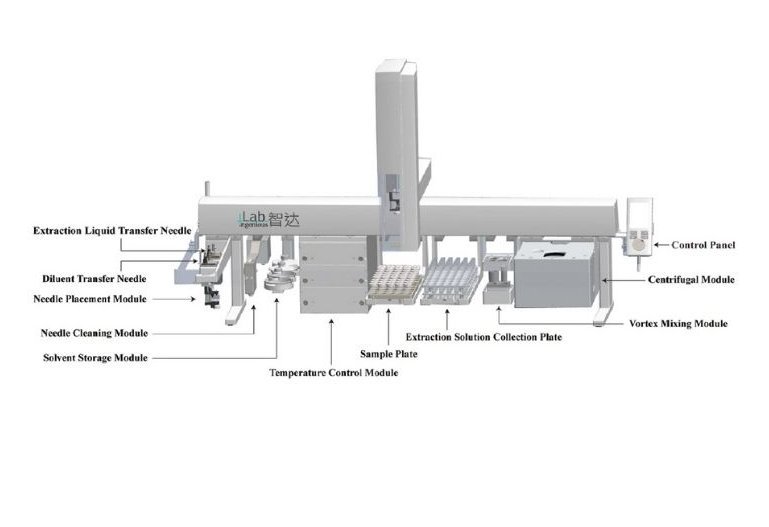
Pushing the boundaries of what's possible, some researchers are developing entirely new automated methods. A remarkable example comes from He et al. (2025), who created a fully automated, high-throughput electro-extraction (EE) platform. Using an advanced dual-headed PAL RTC/RSI system, they developed a workflow that can process 120 samples per day for less than 0.1 Euro per sample. This innovative technique achieves incredible enrichment of target molecules from tiny samples, like 20 microliters of plasma or small mouse muscle tissues, with extraction recoveries up to 99%. This work represents a significant stride towards the future of bioanalysis, showing how novel automation can handle precious, mass-limited samples with exceptional efficiency and precision [7].
Automated Blood Analysis via GC/MS
As Dr. Gernot Poschet, Executive Managing Director of the Metabolomics Core Technology Platform at Heidelberg University, notes, automation is a "clear game-changer" for the field. In a recent interview, he emphasized that "automation of as many steps as possible can highly improve final results" by reducing human error and ensuring data consistency, which is essential for large-scale clinical studies.
This principle is powerfully demonstrated by the work of research consortia. A prime example comes from RECETOX, which developed a fully automated workflow for the metabolomic profiling of blood samples, including the increasingly popular Dried Blood Spots (DBS). Their method, using a robotic platform for direct, in-vial derivatization, achieves excellent reproducibility (CV < 30%) and a high throughput of approximately 40 samples per day. By making these robust, automated methods shareable, consortia like EIRENE benefit from standardized procedures across the field, a critical step towards solving the reproducibility crisis [13].
Read more about their work and open-access methods approach.
This collaborative spirit is shared by leading core facilities like the Metabolomics Core Technology Platform (MCTP) at Heidelberg University, which also relies on automated platforms like the PAL System to support a wide range of research, from basic science to large clinical consortia. By providing access to standardized, high-quality workflows, these facilities are building the foundation for more reliable and impactful science.
These studies point to an exciting and optimistic trend: automation is about more than just speed. It is about fundamentally improving the quality, reliability, and consistency of metabolomic data. By letting robots handle repetitive tasks, we reduce the human introduced variability that can mask true biological discoveries.
This is where the flexible, modular design of modern robotic platforms offers a path forward. Systems like the PAL RTC and PAL RSI are designed as a "Tool Box" for the laboratory [10]. A lab can begin its automation journey with simple liquid injections and, over time, expand its capabilities by adding new modules for more advanced tasks. This modularity empowers laboratories to build custom automated systems that perfectly match their research goals.
The possibilities are vast. A single platform can be configured for headspace analysis of volatile organic compounds (VOCs), or employ dynamic headspace for greater sensitivity. It can perform solid phase microextraction (SPME) with advanced tools like the SPME Arrow, or conduct miniaturized cleanups with Micro SPE cartridges. This flexibility extends across numerous fields, enabling everything from proteomics analysis and automated sample preparation for FAME to critical environmental testing, such as PAH analysis in food or the increasingly important PFAS analysis in drinking water. By solving the challenges of variability and throughput, automation is laying the groundwork for a brighter, more reproducible, and more insightful future in metabolomics and beyond.
References
[1] Cochran, D., Noureldein, M., Bezdeková, D., Schram, A., Howard, R., & Powers, R. (2024). A reproducibility crisis for clinical metabolomics studies. TrAC Trends in Analytical Chemistry, 180, 117918.
[2] Bifarin, O. O., Yelluru, V. S., Simhadri, A., & Fernández, F. M. (2025). A Large Language Model-Powered Map of Metabolomics Research. Analytical Chemistry, 97, 14088-14096.
[3] Siuzdak, G., Activity Metabolomics and Mass Spectrometry; MCC Press: San Diego, USA, 2025. [4] CTC Analytics AG, Pitch Deck PAL System Theme 2025-02. [5] PAL System, Automated, Green, and Efficient: A Compendium on Micromethods in Analytical Sample Preparation, 2025.
[6] Luo, Y.; et al. An automated liquid-liquid extraction platform for high-throughput sample preparation of urinary phthalate metabolites in human biomonitoring. Talanta 2025, 288, 127740.
[7] He, Y.; et al. A fully automated, high-throughput electro-extraction and analysis workflow for acylcarnitines in human plasma and mouse muscle tissues. Analytica Chimica Acta 2025, 1364, 344224.
[8] Mosley, J. D.; et al. Establishing a framework for best practices for quality assurance and quality control in untargeted metabolomics. Metabolomics 2024, 20, 20.
[9] Broeckling, C. D.; et al. Current Practices in LC-MS Untargeted Metabolomics: A Scoping Review on the Use of Pooled Quality Control Samples. Anal. Chem. 2023, 95, 18645–18654.
[10] PAL System, PAL RSI and PAL RTC Sample Prep and Injection Brochure.
[11] Xu, Y., & Goodacre, R. (2025). Mind your Ps and Qs - Caveats in metabolomics data analysis. Trends in Analytical Chemistry, 183, 118064.
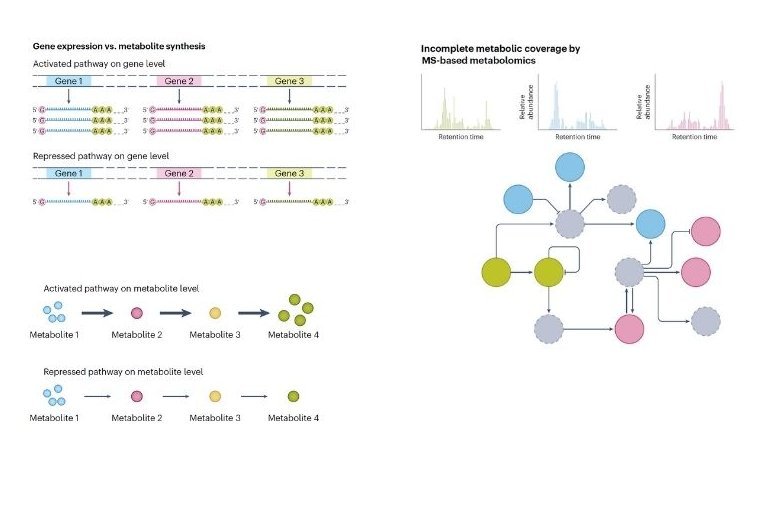
[12] Lee, K. S., Su, X., & Huan, T. (2025). Metabolites are not genes - avoiding the misuse of pathway analysis in metabolomics. Nature Metabolism.
[13] Jbebli, A.; et al. Automated Direct Derivatization and GC/MS Analysis: A Robust Method for Comprehensive Metabolomic Profiling of Dried Blood Spots, Serum, and Plasma. PAL System GC/MS Application Note, Feb 2025.